New BioRxiv pre-print
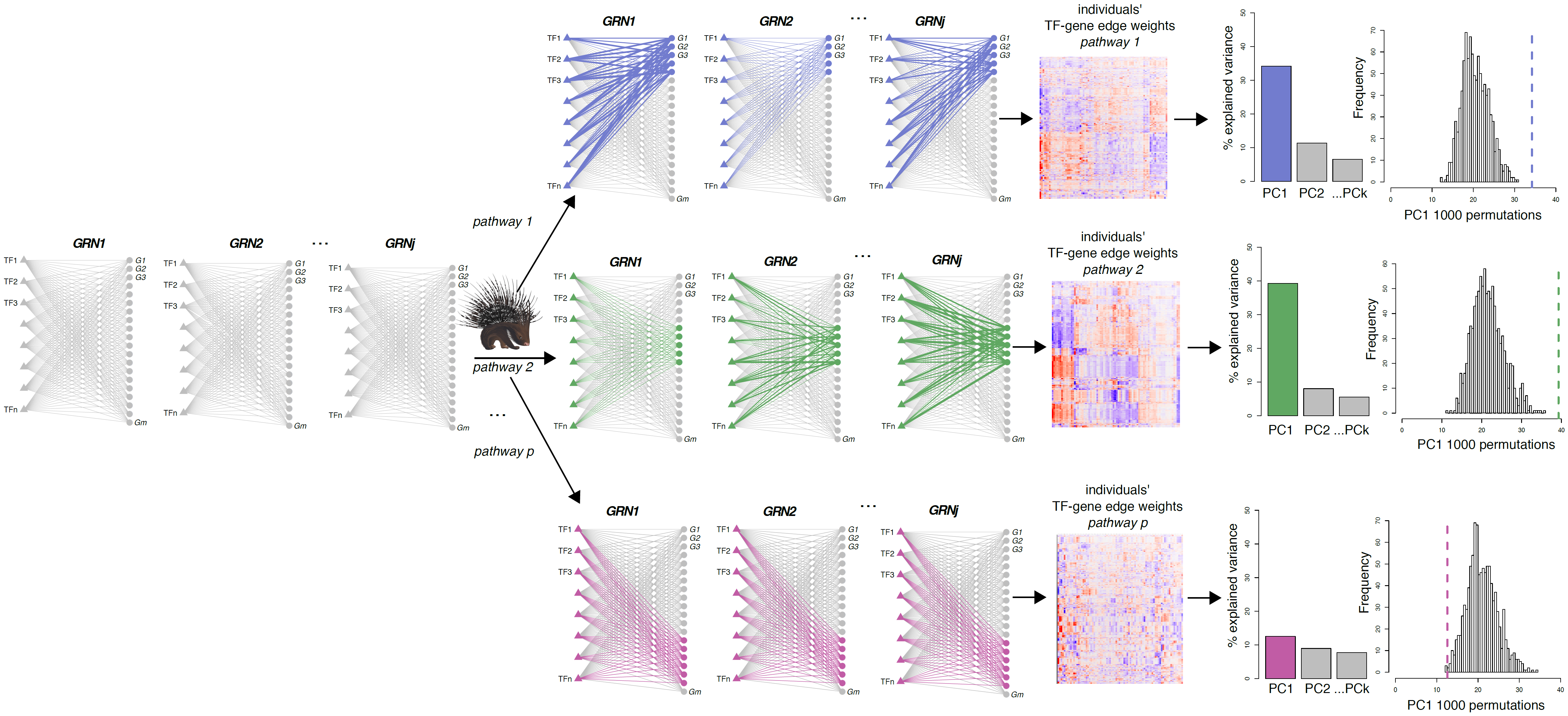
We have a new BioRχiv pre-print, describing our latest tool PORCUPINE and its application to leiomyosarcoma, an aggressive soft-tissue sarcoma subtype. PORCUPINE, which was developed by Tatiana, analyzes complex, genome-wide, patient-specific regulatory networks directly on the network's edge weights. It does this by combining Principal Component Analysis (PCA) with permutations to identify pathways contributing to regulatory heterogeneity in a patient population. Importantly, instead of identifying discrete subtypes, it can map subtle gradients of heterogeneity that may not be detected by standard subtyping approaches.
- Applying PORCUPINE to leiomyosarcoma data, we identified 37 heterogeneously regulated pathways, including targetable pathways such as E2F signaling. These findings were independent of mutations and could thus be a new way of stratifying patients for personalized medicine. The BioRχiv pre-print can be found here. More information on the pre-print can be found here.